Last weekend I took part in Hacktion Potential, a 48-hour computational neuroscience hackathon at the Institut du Cerveau in Paris. Our team worked on decoding mouse maze position from brain signals: mapping high-dimensional neural activity to the (x,y) position of a mouse in a closed environment.
The hackathon goal was to design a decoding method that takes raw hippocampal data (spike activity recorded at ~20 kHz) and predicts spatial location.

Technical story
Since we had very limited time, we kept the approach simple: 1D-CNNs, a few ablations, and a focus on understanding the data and evaluation (splits, window size) rather than pushing state-of-the-art architectures.
Data
Recordings come from the hippocampus with a silicon probe at 20 kHz. Each probe has several shanks recording from different locations. For each decoding timepoint we have the true (x,y) position (sampled at ~15 Hz). Only timepoints where the mouse is moving are used for decoding, since position encoding is physiologically only relevant during movement.
The dataset is built from highpass-filtered, spike-focused activity. For each shank we: detect spikes when voltage exceeds 3× standard deviation on any channel.
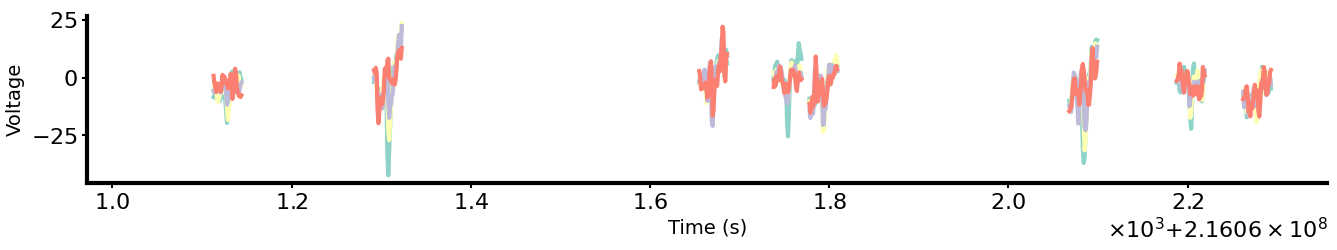
Model architecture and data ablation
We tested whether spike waveforms are needed for decoding. Two models:
- Spiking Embedding Network: only the sequence of spike indices (which shank fired). Trainable embeddings per shank → sequence → 1D-CNN → FC → (x,y).
- Waveform Network: 1D-CNN on the actual waveforms, then global average pooling over the sequence → FC → (x,y).
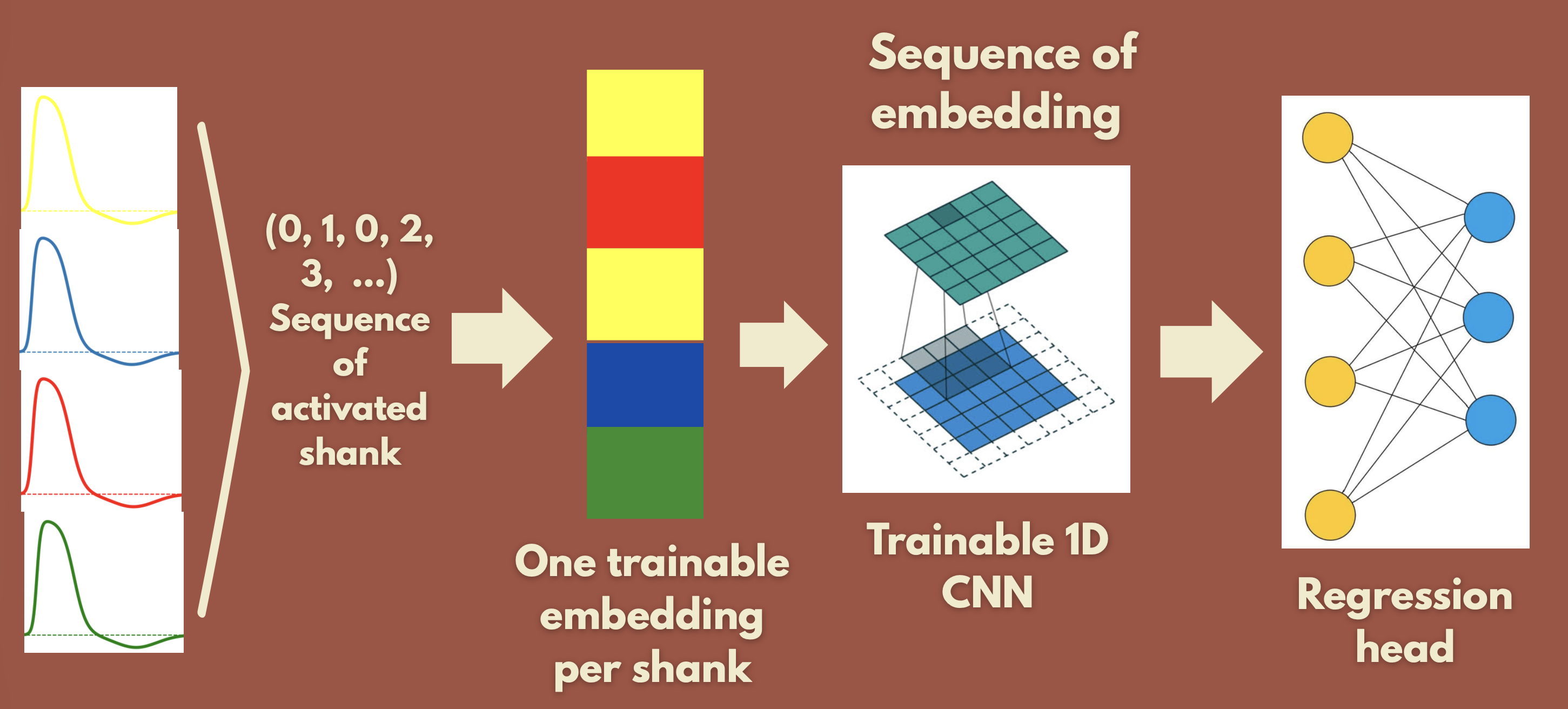
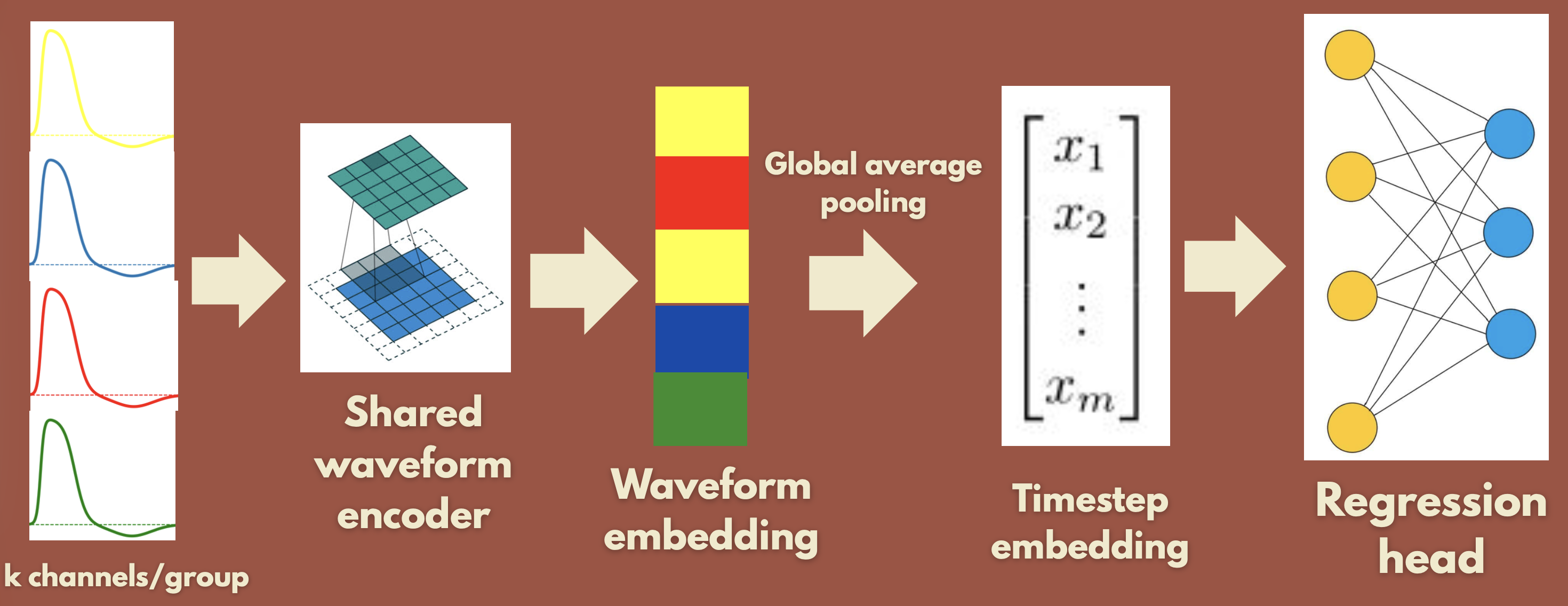


Result: the Spiking Embedding Network does not generalize (low train loss, high test loss). The Waveform Network generalizes well. So waveform information appears necessary, likely because it identifies which neuron fired, not only when. All of this was with a random train/test split.
Window size
We varied the spike window length (18+18, 54+54, 126+126 timesteps around the peak), i.e. 36, 108, and 252 timesteps. All settings learn: 108 converged fastest, consistent with a trade-off between information and noise.
Below we show learning curves for both shuffled and temporal splits (split types are detailed in the next section).






Splitting issues
Train/test split had a large impact. We compared:
- Temporal: train on first 90%, test on last 10% (as suggested).
- Middle: test on middle 10%, train on the rest.
- Shuffled: random split.



| Split type | Lowest train MSE | Lowest test MSE |
|---|---|---|
| Temporal | 0.018 | 0.103 |
| Middle | 0.030 | 0.081 |
| Shuffled | 0.036 | 0.054 |
Here, we did not use a validation set since we did not even tune hyperparameters.
Temporal and middle splits generalize poorly while shuffled seems to generalize better. That suggests that there might be distribution shift or non-stationarity over the session. Another hypothesis is that the data might not carry position information. However, that goes against the results of this study: Zhang, K. et al. (1998) ‘Interpreting Neuronal Population Activity by Reconstruction: Unified Framework With Application to Hippocampal Place Cells’, Journal of Neurophysiology, 79(2), pp. 1017–1044. We might also have some data leakage issues for the shuffled split.

Predictions on the test set are noisy but follow the trajectory. Notice that the model often predicts positions outside the maze.
Future directions
- Model: tune hyperparameters, regularization, data augmentation, cleaner preprocessing to reduce overfitting and shift.
- Generalization: run on multiple mice (current runs are one mouse).
- Multi-subject: train on pooled data across mice.
- Model redesign: condition on maze geometry (e.g. ControlNet-style), predict position as a heatmap (example below), or add past positions as input.

Conclusion
We didn’t win, and the experiments weren’t really conclusive. I still really enjoyed the hackathon. The subject is really interesting and I got to get a glimpse back into neuroscience. I loved the opportunity to work at that pace. The experience was very stimulating.
- Repository: github.com/clementw168/mouse-maze-position-from-brain-signals
- Slides: presentation.pdf in the repo